

This document explains how to reproduce the figures from [1] from the beginning to the end. It describes the following steps:
C++ code of
the dune-tectonic module as well as its dune
dependencies and creating Linux1 executables from it.Each of these steps but the last is optional:
The procedure outlined in the following paragraphs yields an
Lzip-compressed2 tarball
named dune-tectonic.tar.lz. It contains a 2D
executable one-body-problem-2D and a 3D
executable one-body-problem-3D as well as
configuration files, copies of the dynamically linked libraries,
and even a dynamic linker. By downloading such a tarball directly
from
here one can choose
to skip this
section.
The numerical simulations whose results are presented
in [1] are based on
the 2016-PippingKornhuberRosenauOncken branch of
the dune-tectonic
C++ code.
It depends on version 2.5 of a few dune modules (see also the dune.module file) which are available from the DUNE project and the FU Berlin (some from the former, some from the latter).
The building procedure has been automated through docker. In the
following, we will always assume that the
variable tag has been set
to 2016-PippingKornhuberRosenauOncken and that
the git
repository of Dockerfiles has been cloned in the following
way.
tag=2016-PippingKornhuberRosenauOncken
git clone https://git.imp.fu-berlin.de/pipping/${tag}-docker.gitBuilding dune-tectonic is now as simple as:
cd ${tag}-docker
./01-dune-2.5.bash
./02-dune-tectonic.bashDocker makes it easy to automate and sandbox the process of obtaining ready-to-run executables on x86_64 Linux; the entire process could be completed without docker, though. Turning a Dockerfile into a regular shell-script is entirely straightforward. The idea of using docker for reproducible research was put forward in [2]
The purpose of this section is to obtain the following list of
directories, each containing a file
named output.h5:
2d-lab-fpi-tolerance/rfpitol=1e-7
2d-lab-fpi-tolerance/rfpitol=2e-7
2d-lab-fpi-tolerance/rfpitol=3e-7
2d-lab-fpi-tolerance/rfpitol=5e-7
2d-lab-fpi-tolerance/rfpitol=10e-7
2d-lab-fpi-tolerance/rfpitol=20e-7
2d-lab-fpi-tolerance/rfpitol=30e-7
2d-lab-fpi-tolerance/rfpitol=50e-7
2d-lab-fpi-tolerance/rfpitol=100e-7
2d-lab-fpi-tolerance/rfpitol=200e-7
2d-lab-fpi-tolerance/rfpitol=300e-7
2d-lab-fpi-tolerance/rfpitol=500e-7
2d-lab-fpi-tolerance/rfpitol=1000e-7
2d-lab-fpi-tolerance/rfpitol=2000e-7
2d-lab-fpi-tolerance/rfpitol=3000e-7
2d-lab-fpi-tolerance/rfpitol=5000e-7
2d-lab-fpi-tolerance/rfpitol=10000e-7
2d-lab-fpi-tolerance/rfpitol=20000e-7
2d-lab-fpi-tolerance/rfpitol=30000e-7
2d-lab-fpi-tolerance/rfpitol=50000e-7
2d-lab-fpi-tolerance/rfpitol=100000e-7
3d-lab/rtol=1e-5_diam=1e-2These files3 can be downloaded through a script from the aforementioned git repository of Dockerfiles by executing the following commands:
cd ${tag}-docker
./03-fetch-output.bashafter which this entire section can be skipped.
To be able to reproduce all the figures from [1], one needs to run twenty-one 2D computations and a single 3D computation.
Each of the programs below can be passed the
option -io.printProgress true in order to enable a
progress indicator. Without this option, which is off by
default, completing a time step will cause no output to be
written to the console so that it may seem as if the program was
stuck.
The 3D simulation can be run like this (after unpacking the tarball we obtained earlier to the current directory):
## In the directory 3d-lab/rtol=1e-5_diam=1e-2
rtol=1e-5
diam=1e-2
./ld-linux-x86-64.so.2 --library-path . ./one-body-problem-3D \
-v.fpi.tolerance ${rtol} \
-timeSteps.refinementTolerance ${rtol} \
-boundary.friction.smallestDiameter ${diam}while a 2D simulation can be run like this (again, after unpacking the tarball to the current directory):
## In the directory 2d-lab-fpi-tolerance/rfpitol=1e-7
rfpitol=1e-7
./ld-linux-x86-64.so.2 --library-path . ./one-body-problem-2D \
-v.fpi.tolerance ${rfpitol}where rfpitol should take the values
1e-7, 2e-7, 3e-7,
5e-7, 10e-7, 20e-7,
30e-7, 50e-7, 100e-7,
200e-7, 300e-7, 500e-7,
1000e-7, 2000e-7, 3000e-7,
5000e-7, 10000e-7, 20000e-7,
30000e-7, 50000e-7, 100000e-7.
Dependening on the resources available to you, you might want to run all of the 2D simulations in parallel on a cluster (e.g. through slurm), in parallel on one or more multi-core machines (e.g. via GNU Parallel), but probably not in sequence (it would take weeks!). The number of available CPUs is the limiting factor here since none of the 2D simulations needs more than 100M of RAM.
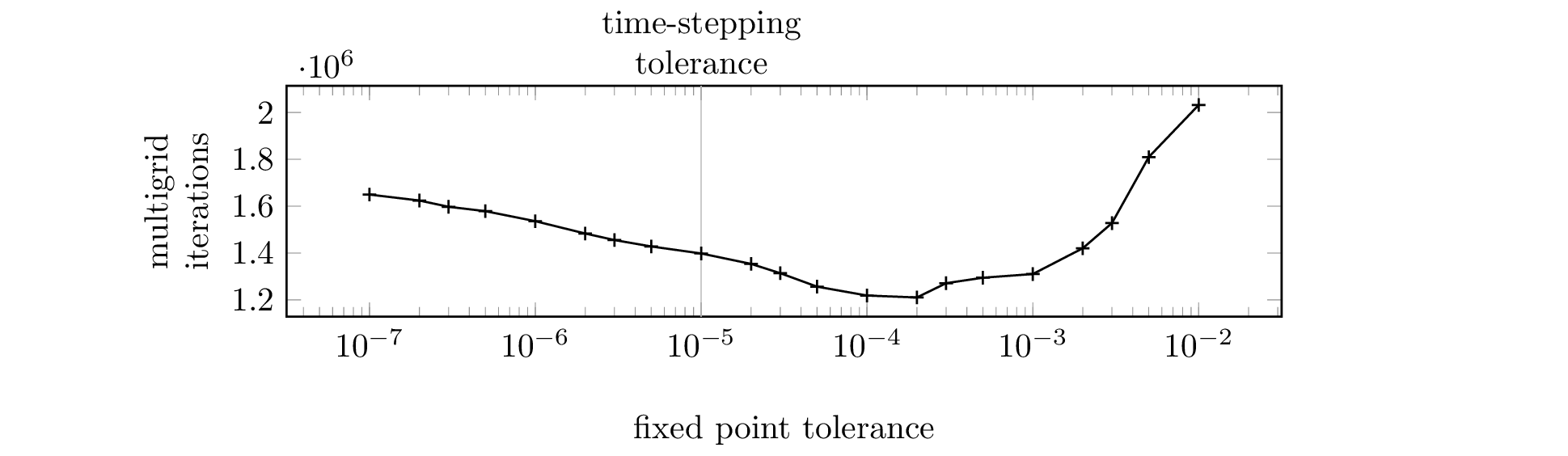
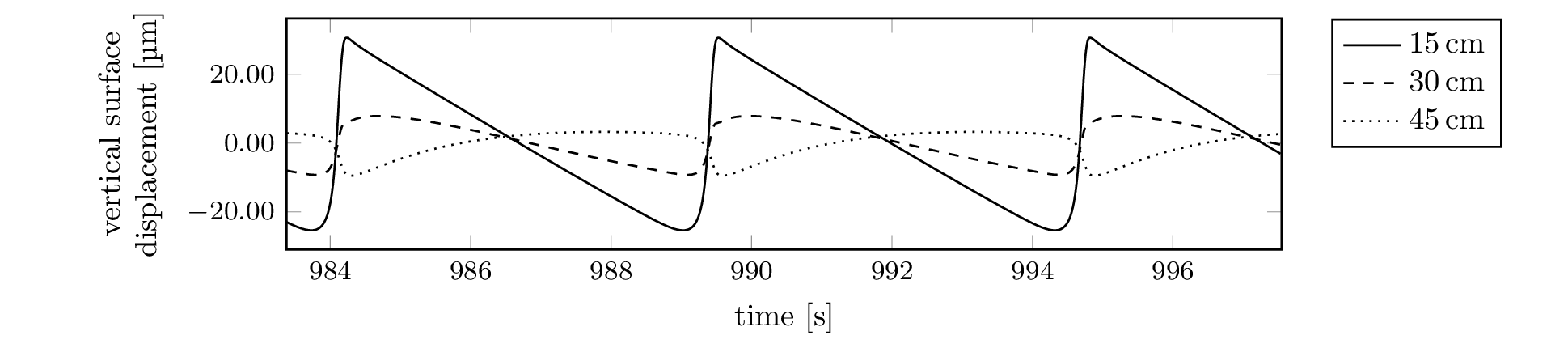
All but one of the 2D simulation plots (Figure 7) only put the
data set
2d-lab-fpi-tolerance/rfpitol=100e-7 to use. If the
reproduction of Figure 7 or the 3D simulation plots is not a
priority, a lot of computing time can be saved by running just this
one simulation.
To be able to talk about seismic events or earthquakes and
quantities related to them, we need to decide on criteria that
allow a machine to determine the spatial and temporal extect of
such an occurrence, and then apply these criteria. This task is
carried out by a collection of R scripts which live
alongside the LaTeX sources we will talk about in the
next section. Once they are run, they populate a directory called
generated with various CSV and contour files. By
downloading the expected contents of this directory as a tarball
from
here one can choose
to skip this
section.
The aforementioned R tools reside in
the tools directory of
the 2016-PippingKornhuberRosenauOncken-plots
git repository.
Before running them, the file config.ini needs to be
edited such that simulation is set to the directory
which contains 2d-lab-fpi-tolerance
and 3d-lab from the previous section as immediate
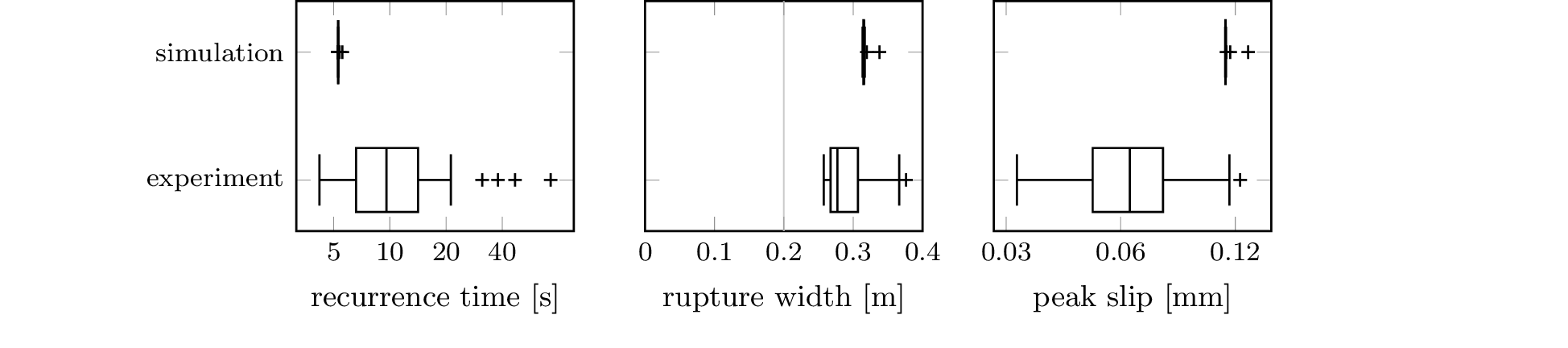
subdirectories. The boxplot from Figure 5 also requires a local
copy of the file B_Scale-model-earthquake-data.zip
from [3], whose location
should be recorded in config.ini as
the experiment directory.
Finally, we install the necessary R packages (namely
Rcpp and h5) and run the scripts via
./tools/generate-2d.bash
./tools/generate-3d.bash
./tools/generate-others.bashThis entire process, too, is automated through a Dockerfile, so
that one only needs to take the following steps to obtain the
file generated.tar.lz.
cd ${tag}-docker
./04-R-tools.bashThe LaTeX/pgfplots
code that is used to produce the final figures is available from
the 2016-PippingKornhuberRosenauOncken-plots
git repository that also hosts the R tools from the
previous section which we used to populate the directory
called generated. We will now need the contents of
this directory.
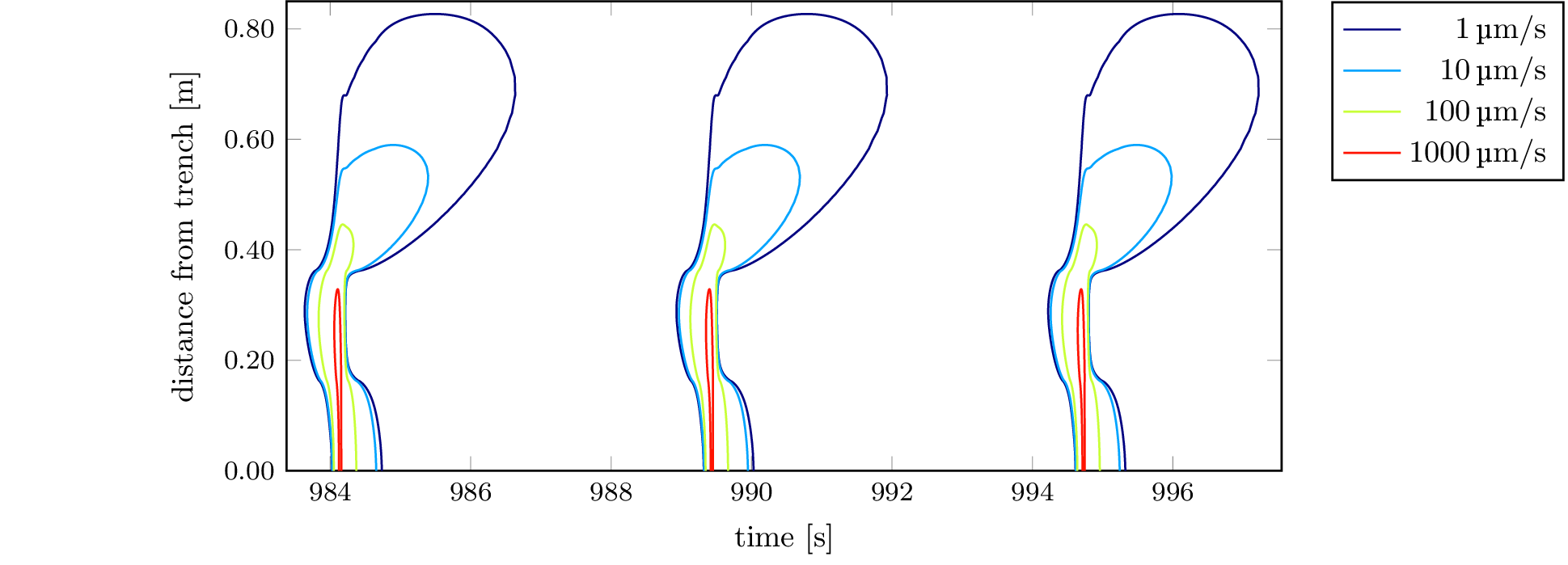
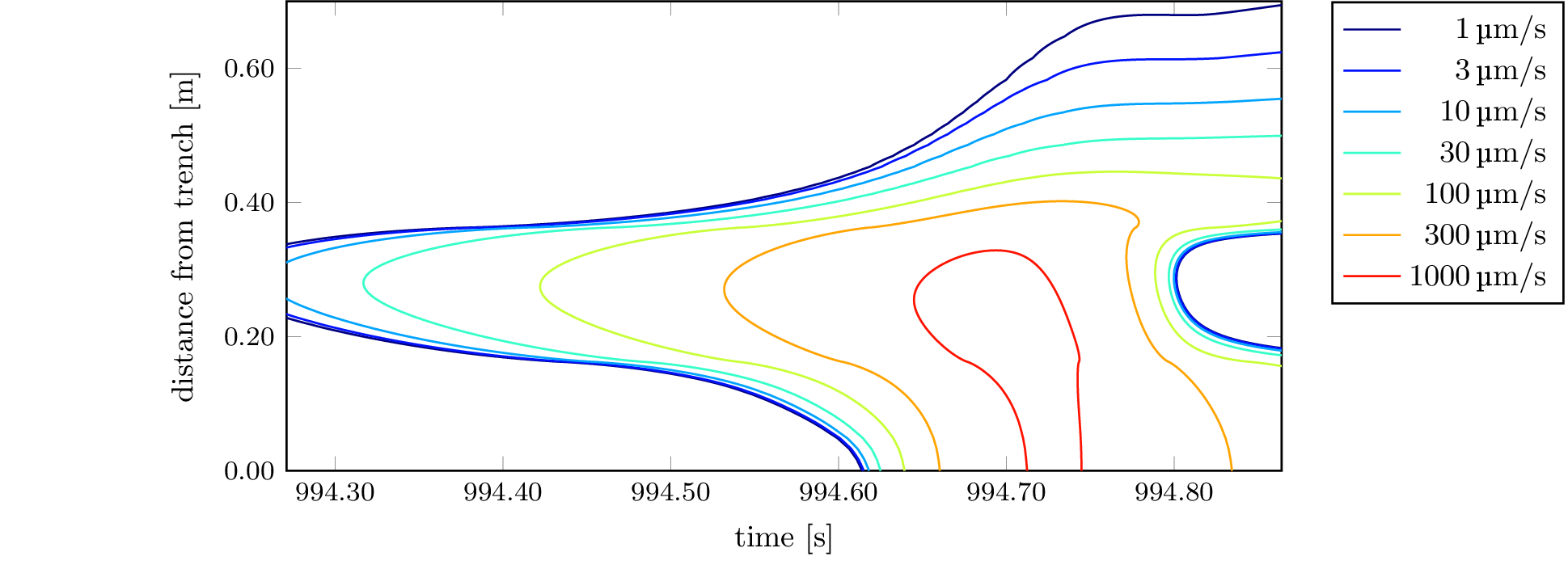
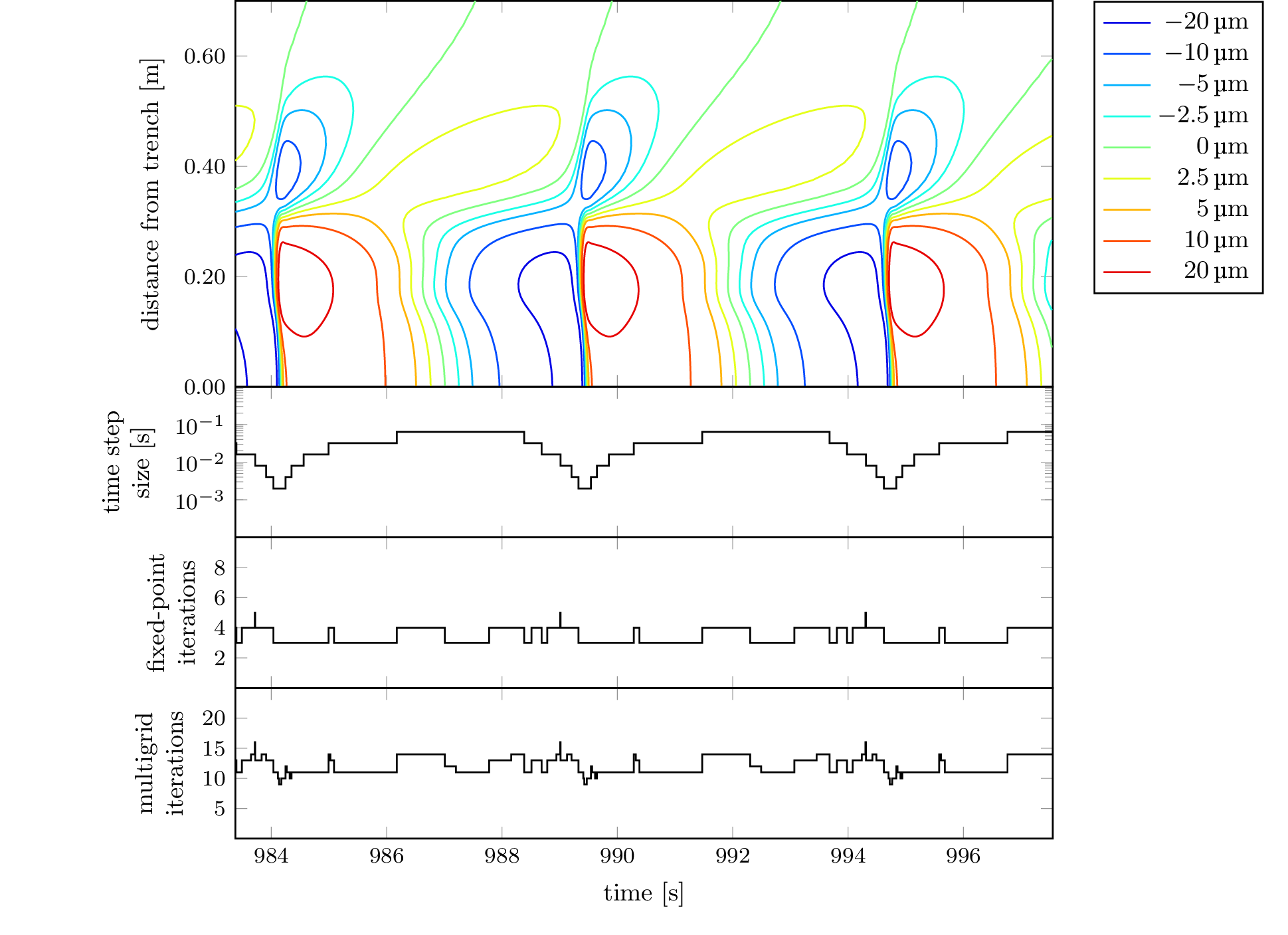
To recreate the plot from Figure 6, which is contained
in 2d-dip-contours-performance.tikz, we can use the
standalone LaTeX
document standalone/2d-dip-contours-performance.tex
which contains nothing but this figure and process it with a LaTeX
processor, e.g. via
latexmk -pdf standalone/2d-dip-contours-performance.texThe process of doing this for all figures can also be automated via
make -f standalone/Makefile pdfIn either case, the resulting PDF files can be found in the current working directory. The Makefile also makes it easy to turn these PDF files into PNG files via
make -f standalone/Makefile pngassuming that ImageMagick is installed.
Naturally, also these steps are automated through a Dockerfile,
so that one only needs to run the following commands to obtain the
file figures.tar.lz (filled with PNG images), which
is also available from here):
cd ${tag}-docker
./05-latex-figures.bashThe above steps should generate a series of PNG images which are shown here.








E. Pipping, R. Kornhuber, M. Rosenau, and O. Oncken: On the efficient and reliable numerical solution of rate-and-state friction problems, Geophys. J. Int. 204.3 (2016) 1858–1866. DOI
C. Boettiger: An introduction to Docker for reproducible research, with examples from the R environment, ACM SIGOPS OSR 49.1. (2015) 71–79. DOI arXiv
M. Rosenau, F. Corbi, S. Dominguez, M. Rudolf, M. Ritter, E. Pipping: Supplement to "Analogue earthquakes and seismic cycles: Experimental modelling across timescales", GFZ Data Services (2016). DOI
Creating a macOS executable works just as well, prebuilt binaries are not provided, however. The same goes for Linux on architectures other than x86_64.↩
Lzip was used here because it led to considerably higher rates of compression than the more widely available gzip, is less susceptible to corruption and does not contain the modification time of the input so that identical input will lead to identical output. It is hoped that Lzip will see more widespread use in the future so that this choice will lead to less and less discomfort for the user.↩
The files you can download here are different from the ones that
one-body-problem-2D generates in that they've
been post-processed as follows:
h5repack -l CONTI output.h5 repacked.h5
h5repack -f GZIP=9 repacked.h5 compressed.h5The purpose is quicker random access and a smaller file size.↩